Machine learning allows researchers to see across the spectrum
[ad_1]
(Nanowerk News) Organic chemistry, the study of carbon-based molecules, underlies not only the science of living organisms, but is essential for many current and future technologies, such as organic light-emitting diode (OLED) displays. Understanding the molecular electronic structure of a material is the key to predicting the chemical properties of that material.
In a recently published study (Journal of Physical Chemistry Letters, “Ground-state Electronic Structure Prediction from Core-Loss Spectra of Organic Molecules by Machine Learning”) by researchers at the Institute of Industrial Science, The University of Tokyo, a machine learning algorithm was developed to predict the density of states in organic molecules, that is, the number of energy levels that electrons can occupy in the ground state in a molecule of material. These predictions, based on spectral data, can greatly assist organic chemists and materials scientists when analyzing carbon-based molecules.
Experimental techniques often used to find the density of states can be difficult to interpret. This is especially true for the method known as nuclear loss spectroscopy, which combines energy loss near edge spectroscopy (ELNES) and structure near edge X-ray absorption (XANES). This method shines a beam of electrons or X-rays on a sample of the material; the resulting scattering of electrons and the measurement of the energy emitted by the molecules of the material allows the density of the desired molecular state to be measured. However, the information that the spectrum has is only in the absence (empty) electron states of the excited molecule.
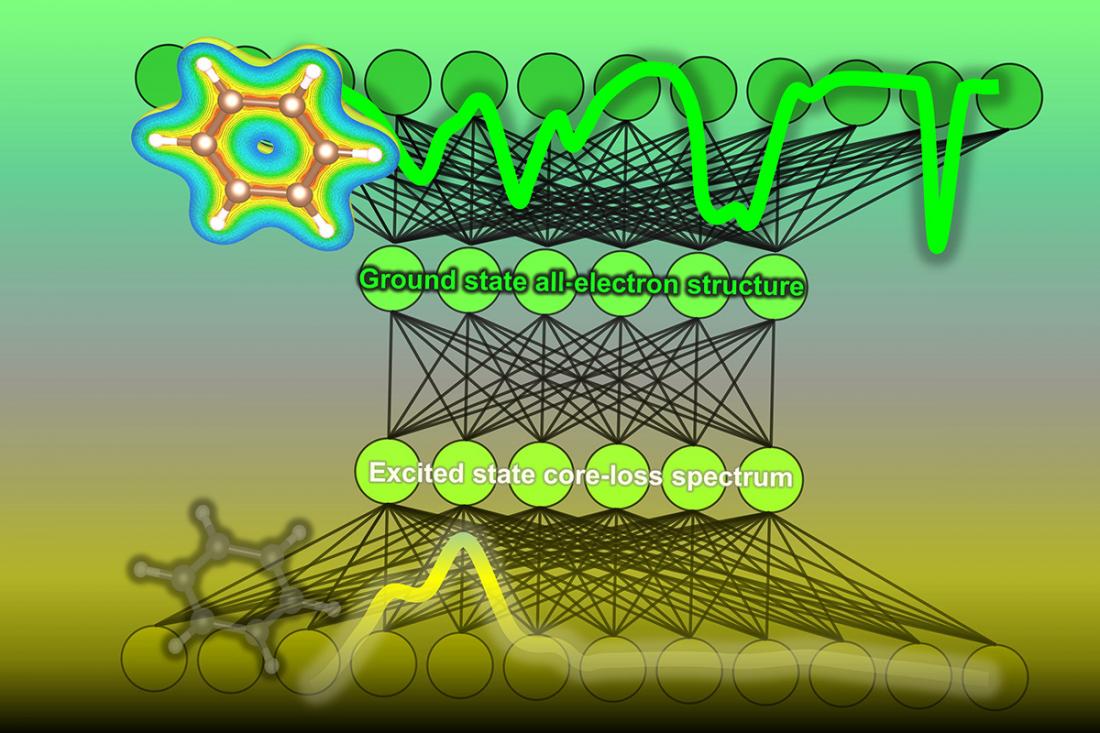
To solve this problem, a team at the Institute of Industrial Science, The University of Tokyo, trained a machine learning model of a neural network to analyze core loss spectroscopy data and predict the density of electronic states. First, a database was constructed by calculating the density of states and the corresponding core loss spectra for more than 22,000 molecules. They also added some simulated sounds. Then, the algorithm is trained on a core-loss spectrum and optimized to predict the correct state densities of occupied and unfilled ground states.
“We are trying to extrapolate the predictions to larger molecules using a model trained by smaller molecules. We found that accuracy can be improved by excluding small molecules,” explains lead author Po-Yen Chen.
The team also found that by using smoothing preprocessing and adding specific noise to the data, state density predictions could be improved, which could accelerate the adoption of predictive models for use on real data.
“Our work can help researchers understand molecular material properties and accelerate the design of functional molecules,” said senior author Teruyasu Mizoguchi. This can include drugs and other interesting compounds.
[ad_2]
Source link